The miRNA Enrichment Analysis and Annotation Tool (miEAA) facilitates the functional analysis of sets of miRNAs. With
the increase in the amount of data regarding miRNAs, there is also an increase in the need for tools to help
analyzing them.
MiEAA is one of the tools in this regard. It is based on GeneTrail, which is an enrichment analysis tool for
gene sets. In contrast to GeneTrail, miEAA is tailored for miRNA precursors and mature miRNAs of multiple frequently investigated species, for example human, mice, or rat.
- Over-representation analysis (ORA)
- Gene set enrichment analysis (GSEA) adapted for microRNAs
- Different category sets for mature miRNAs and precursors
- Broad range of incorporated databases for a total of 10 species,
comprising H. sapiens, M. musculus, R. norvegicus, A. thaliana, B. taurus, C. elegans, D. melanogaster, D. rerio, G. gallus, and S. scrofa - More than 130,000 categories available for analysis
- MiEAA features IDs from current miRBase release v22
- Down-stream visualization of results, e.g. miRNA to category heatmaps
- Easy remote-access to conduct enrichment analyses using either browsable Web-API or python / R
Adapted from the latest publication:
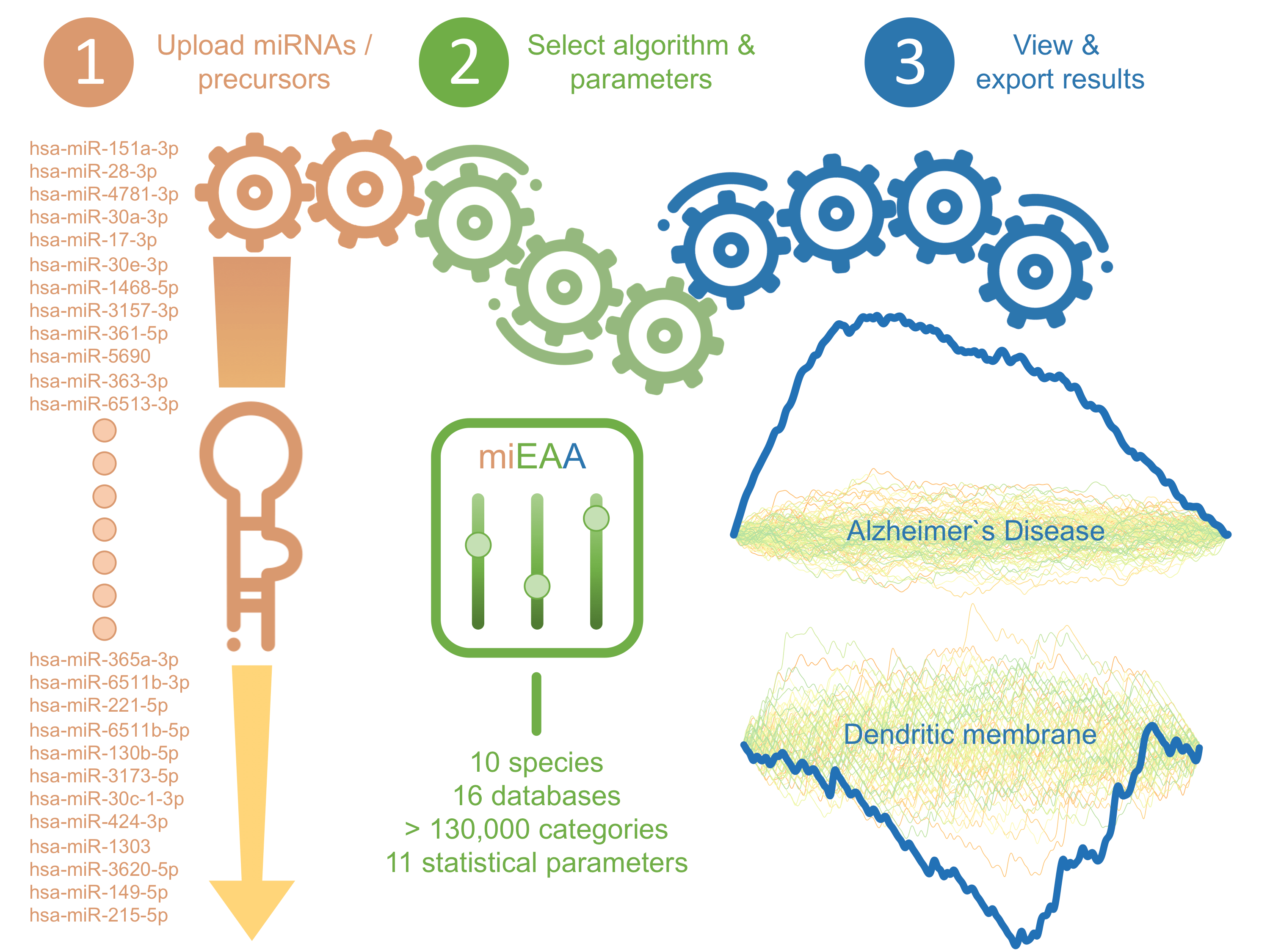
miEAA 2.0 offers over-/under-representation analysis and gene set enrichment analysis for 10 species. The main steps are: 1) upload of a list of miRNAs or precursors, 2) selection of the desired algorithm and all statistical parameters, and 3) the visualization of results in interactive elements.
Adapted from the original publication:
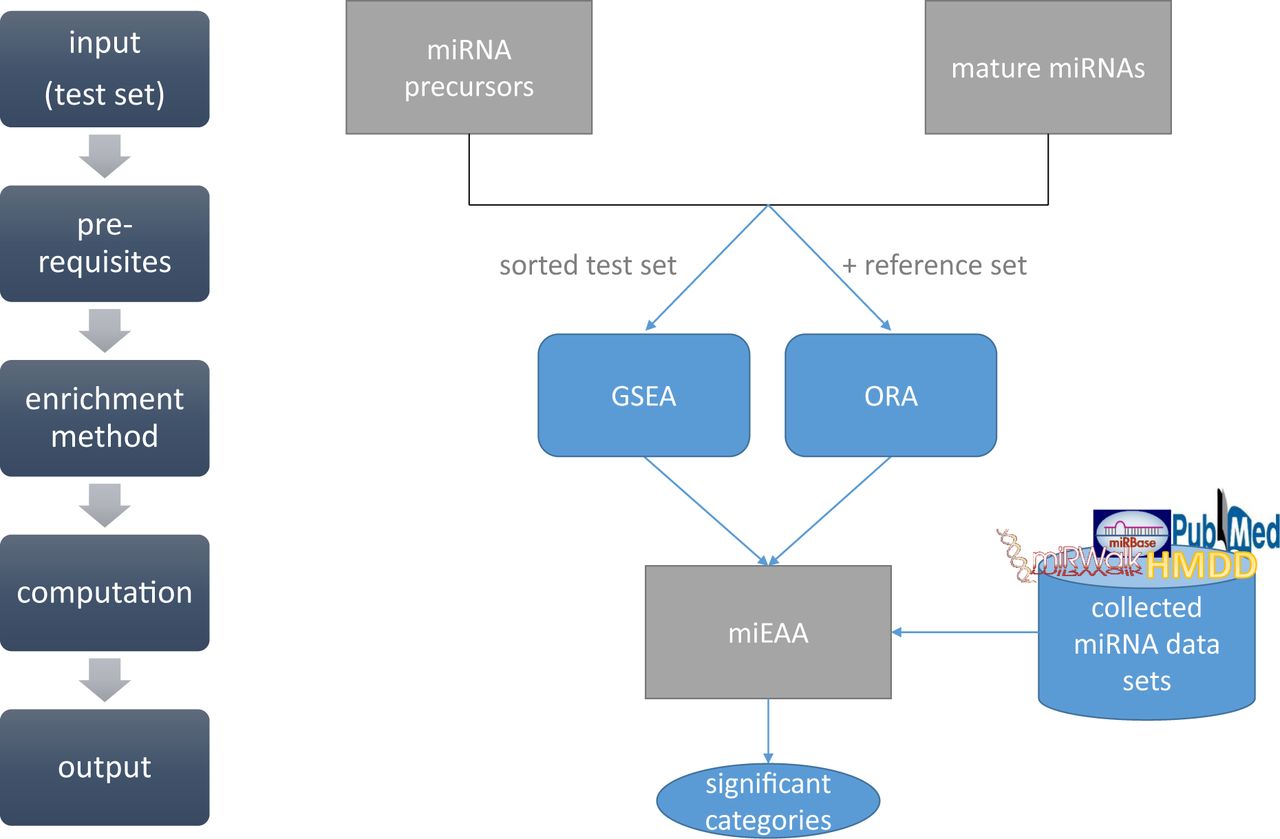
Workflow of miEAA. Input for miEAA are either miRNA or precursor names from miRBase [...]. After uploading a plain text file containing these names, the user can chose the enrichment method: ORA or GSEA.
Depending on this choice, the user can provide an own reference set for ORA. For GSEA, the input list must be sorted by a meaningful criterion.
In a last step, the user can choose the categories that should be analyzed, as well as the P-value significance threshold and adjustment method.
After computation, the results are presented in a tabular format on an HTML web site and can also be downloaded as Excel sheet or tab-separated text file.
If you use our web service, please cite the corresponding publication:
Aparicio-Puerta et al.
miEAA 2023: updates, new functional microRNA sets and improved enrichment visualizations



